In Magnetic Resonance Spectroscopy (MRS), spectra are initially reconstructed in the frequency domain, measured in Hertz (Hz). This relative frequency axis () is strictly dependent on the hardware—specifically, the strength of the main magnetic field ().
To compare spectra across different scanners (e.g., 1.5T vs 3T vs 7T), we convert this hardware-dependent axis into a hardware-independent axis called chemical shift, measured in parts-per-million (ppm).
xmris provides dedicated methods to swap between these coordinate systems seamlessly while tracking the physical metadata.
The Physics & Math¶
The conversion relies on two critical pieces of metadata that must be stored in your xarray.DataArray.attrs:
reference_frequency: The Larmor frequency of the target nucleus (in MHz).carrier_ppm: The absolute chemical shift located exactly at the center of your acquisition band (0 Hz). For H MRS, this is typically the water resonance at 4.7 ppm.
The mathematical relationships are defined as:
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
# Ensure the accessor is registered
import xmris1. Generating a Synthetic Spectrum¶
Let’s generate a mock frequency-domain spectrum (in Hz) acquired on a 3T scanner. We must stamp the required physics metadata into the .attrs.
# Simulated acquisition parameters for 3T
mhz = 127.7 # 1H resonance frequency at 3T
carrier = 4.7 # Transmitter centered on water
n_points = 1024
sw_hz = 2000.0 # 2 kHz spectral width
# Generate relative Hz coordinates centered at 0
hz_coords = np.linspace(-sw_hz/2, sw_hz/2, n_points)
# Simulate two peaks: Water (at 0 Hz / 4.7 ppm) and NAA (at roughly -345 Hz / 2.0 ppm)
water_peak = 1.0 / (1 + ((hz_coords - 0) / 10)**2)
naa_peak = 0.5 / (1 + ((hz_coords - (-345)) / 8)**2)
spec_data = water_peak + naa_peak
# Xarray construction with strictly required attrs
da_freq = xr.DataArray(
spec_data,
dims=["frequency"],
coords={"frequency": hz_coords},
attrs={
"reference_frequency": mhz,
"carrier_ppm": carrier,
"sequence": "PRESS"
}
)
fig, ax = plt.subplots(figsize=(8, 3))
da_freq.plot(ax=ax)
ax.set_title("Spectrum in Relative Frequency (Hz)")
ax.set_xlabel("Frequency [Hz]")
# Note: In MRS, frequency/ppm axes are traditionally plotted right-to-left
ax.invert_xaxis()
plt.show()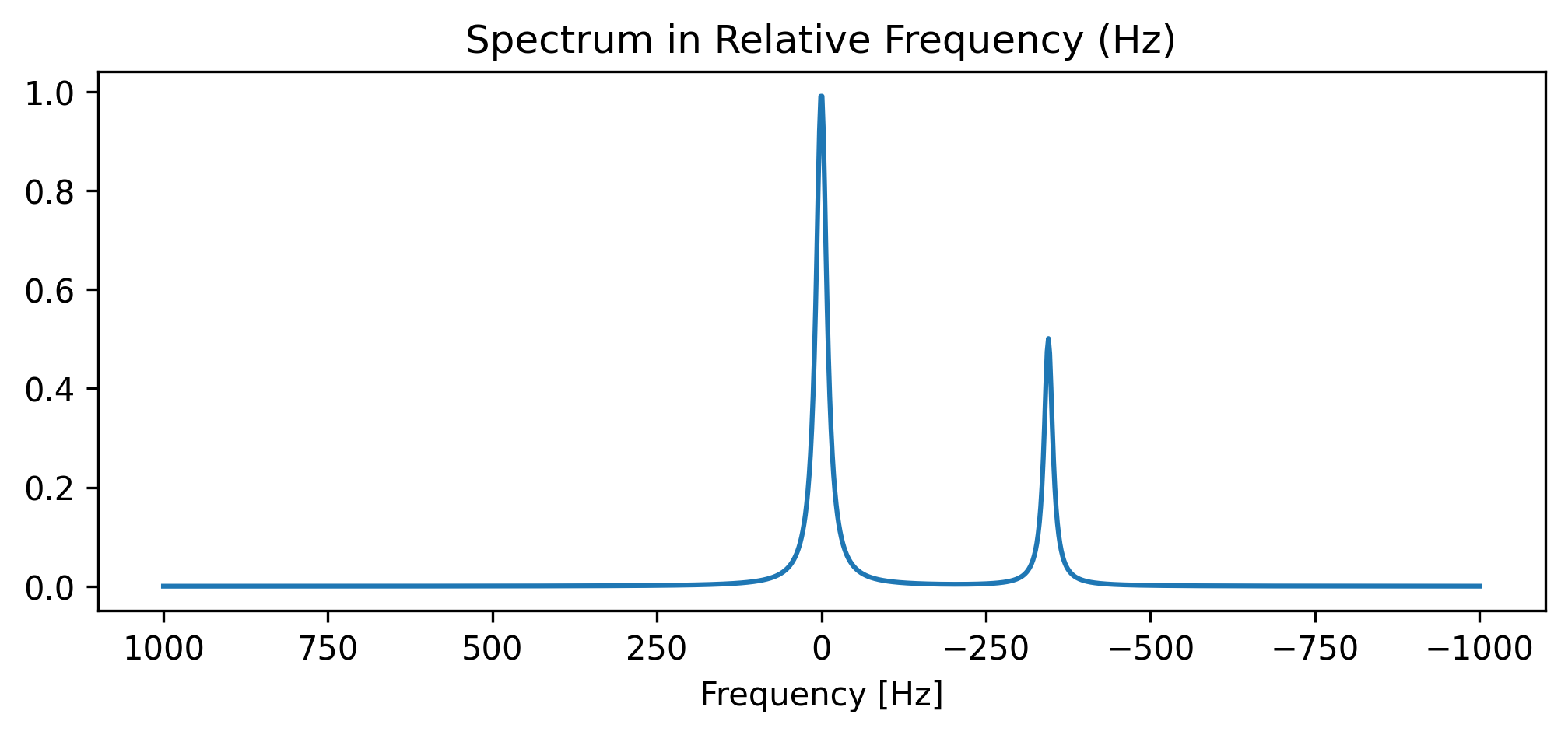
2. Converting to Chemical Shift (ppm)¶
Using .xmr.to_ppm(), xmris calculates the new chemical shift coordinates, creates a properly labeled axis, and automatically swaps the primary dimension of the DataArray from "frequency" to "chemical_shift".
# Convert to ppm using the dimension name as a plain string
da_ppm = da_freq.xmr.to_ppm(dim="frequency")
fig, ax = plt.subplots(figsize=(8, 3))
da_ppm.plot(ax=ax, color="tab:green")
ax.set_title("Spectrum in Absolute Chemical Shift (ppm)")
ax.set_xlabel("Chemical Shift [ppm]")
ax.invert_xaxis()
plt.show()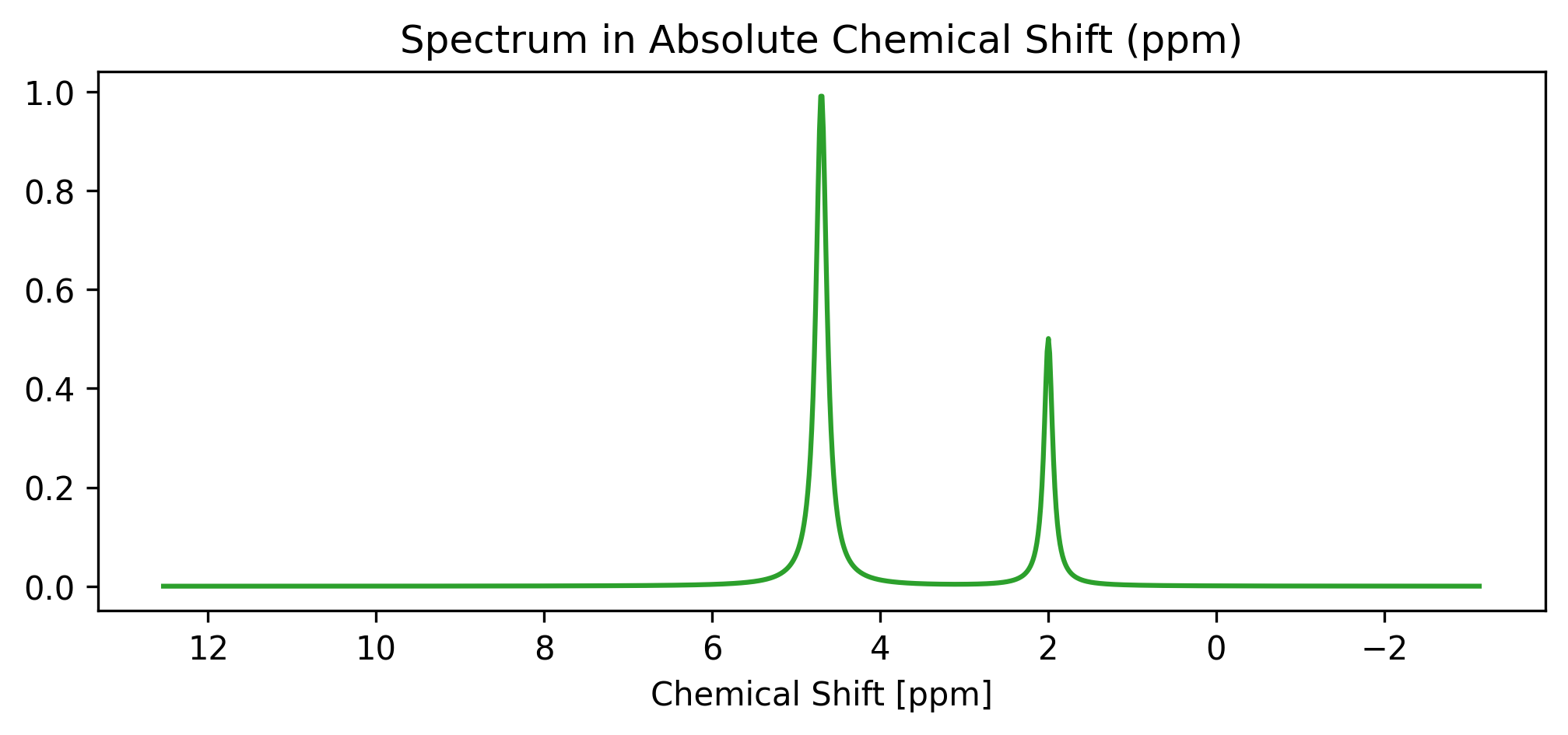
Notice how the water peak is now correctly centered at 4.7 ppm, and the NAA peak is positioned around 2.0 ppm.
Under the Hood: The @requires_attrs Decorator
If you attempt to call to_ppm() on an array that is missing the "reference_frequency" or "carrier_ppm" attributes, xmris will instantly throw a helpful error before any math is executed.
This strict validation guarantees that your physical conversions are always rooted in reality, and prevents silent mathematical drift!
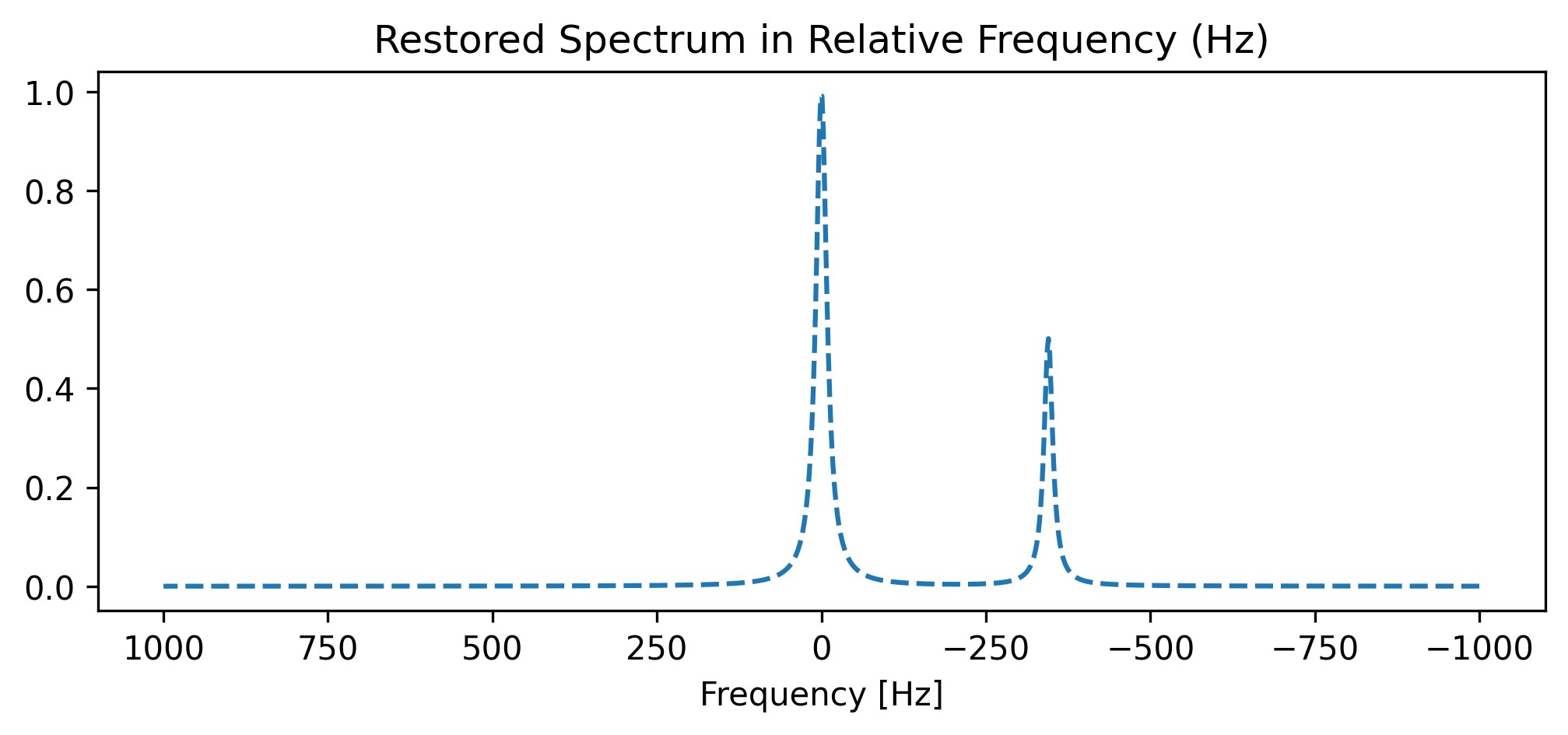
3. Converting back to Frequency (Hz)¶
The operation is perfectly invertible using .xmr.to_hz().
Because xmris preserves data lineage, the metadata attributes are completely untouched during the first transformation, meaning the DataArray has everything it needs to reconstruct the original Hz axis.
# Convert back to relative frequency (Hz)
da_hz_restored = da_ppm.xmr.to_hz(dim="chemical_shift")
fig, ax = plt.subplots(figsize=(8, 3))
da_hz_restored.plot(ax=ax, color="tab:blue", linestyle="--")
ax.set_title("Restored Spectrum in Relative Frequency (Hz)")
ax.set_xlabel("Frequency [Hz]")
ax.invert_xaxis()
plt.show()