Manual phase correction is often tedious. The .xmr.autophase() method automates this process by utilizing global optimization algorithms to find the ideal zero-order () and first-order () phase angles.
Because Magnetic Resonance Spectra vary wildly in density and Signal-to-Noise Ratio (SNR), xmris provides three distinct scoring methods:
acme: Minimizes spectral entropy. Excellent for dense, high-SNR spectra with multiple overlapping peaks.positivity: Maximizes positive real signal and heavily penalizes negative signal within a defined Region of Interest (ROI). Excellent for sparse, noisy spectra.peak_minima: Minimizes the difference between the minima on either side of a target peak. Good for isolated peaks on a rolling baseline.
Imports & plotting function to compare spectra
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
# Import xmris simulation tools and ensure accessors are registered
import xmris
from xmris.fitting.simulation import simulate_fid
def plot_spectra(spectra_list, title, target_coord=None, peak_width=None, xlim=None):
"""Dynamically plots an arbitrary number of spectra for visual inspection."""
n_plots = len(spectra_list)
fig, axes = plt.subplots(n_plots, 1, figsize=(10, 2.2 * n_plots), sharex=True)
if n_plots == 1:
axes = [axes]
fig.suptitle(title, fontsize=14, fontweight="bold")
for ax, (da, label) in zip(axes, spectra_list):
if "Distorted" in label:
ax.plot(da.coords["chemical_shift"], np.real(da.values), color="gray", lw=2)
elif "Corrected" in label:
ax.plot(da.coords["chemical_shift"], np.real(da.values), color="blue", lw=2)
else:
ax.plot(da.coords["chemical_shift"], np.real(da.values), color="black", lw=1.2)
ax.set_ylabel("Amplitude")
ax.legend([label], loc="upper right")
ax.grid(True, alpha=0.3)
ax.axhline(0, color="red", linestyle="--", alpha=0.5)
if target_coord is not None and peak_width is not None:
roi_start = target_coord + (peak_width / 2.0)
roi_end = target_coord - (peak_width / 2.0)
ax.axvspan(roi_start, roi_end, color='green', alpha=0.1)
if xlim is not None:
ax.set_xlim(*xlim)
axes[-1].set_xlabel("Chemical Shift (ppm)")
axes[-1].invert_xaxis()
plt.tight_layout()
plt.show()1. Complex Multi-Peak Spectra (ACME)¶
The ACME algorithm works best when there is plenty of signal to evaluate across the entire spectrum. Here, we simulate a dense spectrum, intentionally ruin the phase, and compare the automated solvers.
As expected, acme reconstructs the ground truth perfectly, whereas local ROI solvers struggle because they ignore the wider spectral context.
Make a dummy 1H spectrum
# 1. Simulate an ideal, perfectly phased FID and convert to spectrum
fid_ideal = simulate_fid(
amplitudes=[100, 60, 40, 20], chemical_shifts=[2.0, 3.0, 3.2, 1.3],
reference_frequency=123.2, carrier_ppm=0.0, dampings=[30, 25, 25, 40],
dead_time=0.0, phases=0.0, target_snr=50, n_points=2048
)
spec_ideal = fid_ideal.xmr.to_spectrum().xmr.to_ppm()
# 2. Distort it natively in the frequency domain
p0_distort, p1_distort, pivot_val = 60.0, -800.0, 0.0
spec_distorted = spec_ideal.xmr.phase(
dim="chemical_shift", p0=p0_distort, p1=p1_distort, pivot=pivot_val
)
# 3. Apply manual correction (double apply with inverted signs)
spec_corrected = spec_distorted.xmr.phase(
dim="chemical_shift", p0=-p0_distort, p1=-p1_distort, pivot=pivot_val
)# Step 1-3.
# --> Simulation of the spectrum is hidden in the collpased cell above.
# 4. Run Autophasing
acme_res = spec_distorted.xmr.autophase(method="acme", dim="chemical_shift")
pkmin_res = spec_distorted.xmr.autophase(method="peak_minima", peak_width=5.0, dim="chemical_shift")
posit_res = spec_distorted.xmr.autophase(method="positivity", peak_width=5.0, dim="chemical_shift")
# 5. Plot
plot_spectra([
(spec_distorted, "Distorted Spectrum (xmr.phase)"),
(spec_corrected, "Corrected Spectrum (Manual Inverse)"),
(acme_res, "ACME Phased"),
(pkmin_res, "Peak Minima Phased"),
(posit_res, "Positivity Phased")
], "TEST 1: Dense Spectrum (High SNR)", xlim=(-4, 9))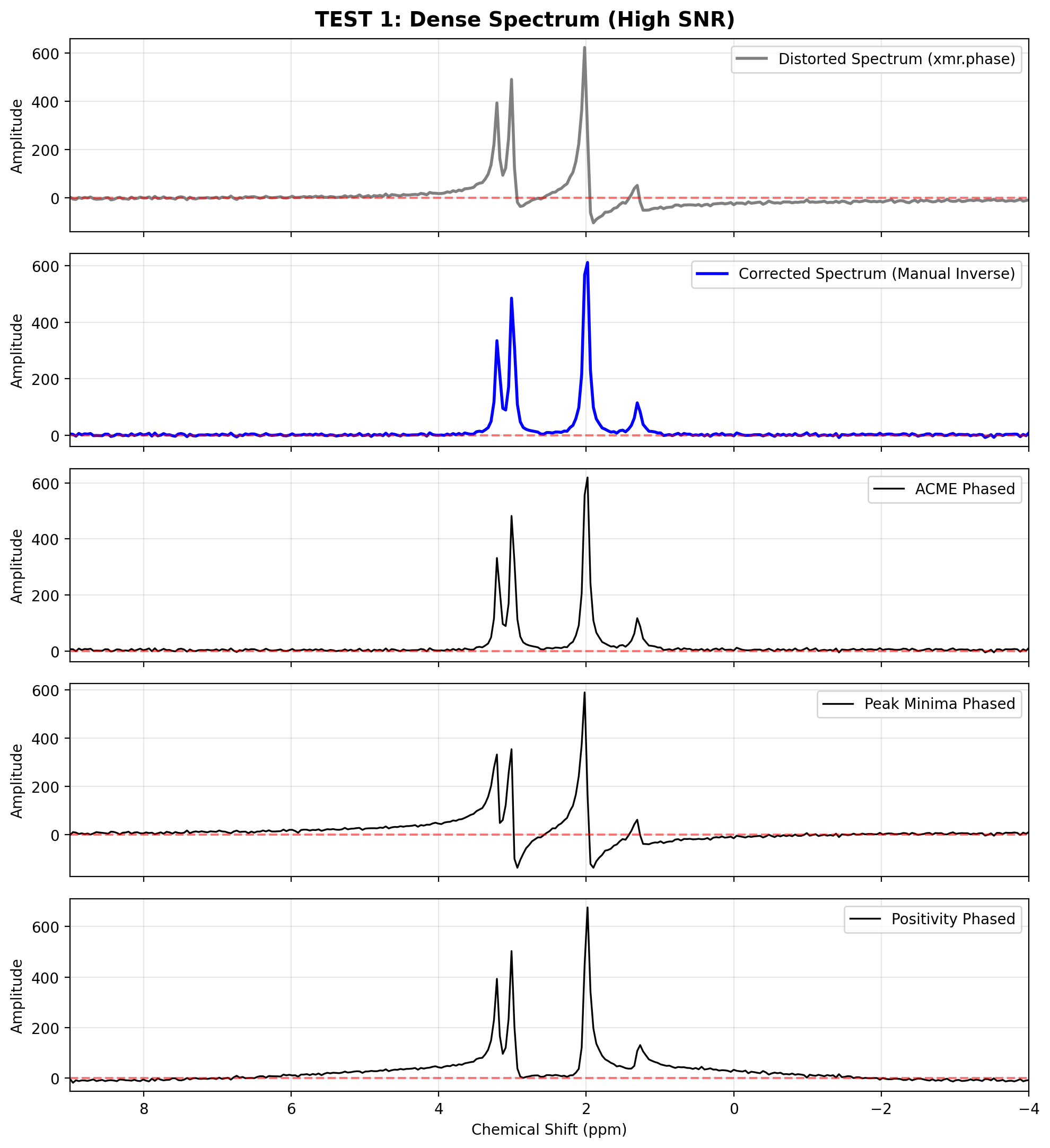
2. Sparse, Noisy Spectra (Positivity)¶
When dealing with sparse spectra (e.g., hyperpolarized or isolated peaks) hidden in high noise, entropy-based methods like ACME often fail because the noise dominates the objective function.
In these cases, utilizing the positivity or peak_minima methods over a localized peak_width around the highest signal is significantly more robust.
Make a dummy 13C spectrum
# 1. Simulate an ideal, but NOISY FID
fid_noisy = simulate_fid(
amplitudes=[100, 20], chemical_shifts=[171.0, 183.0],
reference_frequency=32.1, carrier_ppm=175.0, spectral_width=5000,
dampings=[15, 15], dead_time=0.0, phases=0.0, target_snr=4, n_points=1024
)
# Exponential apodization helps smooth the noise floor
spec_noisy = fid_noisy.xmr.apodize_exp(lb=10.0).xmr.to_spectrum().xmr.to_ppm()
# 2. Distort it
p0_distort_s, p1_distort_s, pivot_s = -45.0, 500.0, 175.0
spec_distorted_s = spec_noisy.xmr.phase(
dim="chemical_shift", p0=p0_distort_s, p1=p1_distort_s, pivot=pivot_s
)
# 3. Manual correction
spec_corrected_s = spec_distorted_s.xmr.phase(
dim="chemical_shift", p0=-p0_distort_s, p1=-p1_distort_s, pivot=pivot_s
)# Step 1-3.
# --> Simulation of the spectrum is hidden in the collpased cell above.
# 4. Autophase
acme_s = spec_distorted_s.xmr.autophase(dim="chemical_shift", method="acme")
pkmin_s = spec_distorted_s.xmr.autophase(dim="chemical_shift", method="peak_minima", peak_width=10.0)
posit_s = spec_distorted_s.xmr.autophase(dim="chemical_shift", method="positivity", peak_width=10.0)
# 5. Plot
plot_spectra([
(spec_distorted_s, "Distorted Spectrum (xmr.phase)"),
(spec_corrected_s, "Corrected Spectrum (Manual Inverse)"),
(acme_s, "ACME Phased"),
(pkmin_s, "Peak Minima Phased"),
(posit_s, "Positivity Phased")
], "TEST 2: Sparse Spectrum (Low SNR)")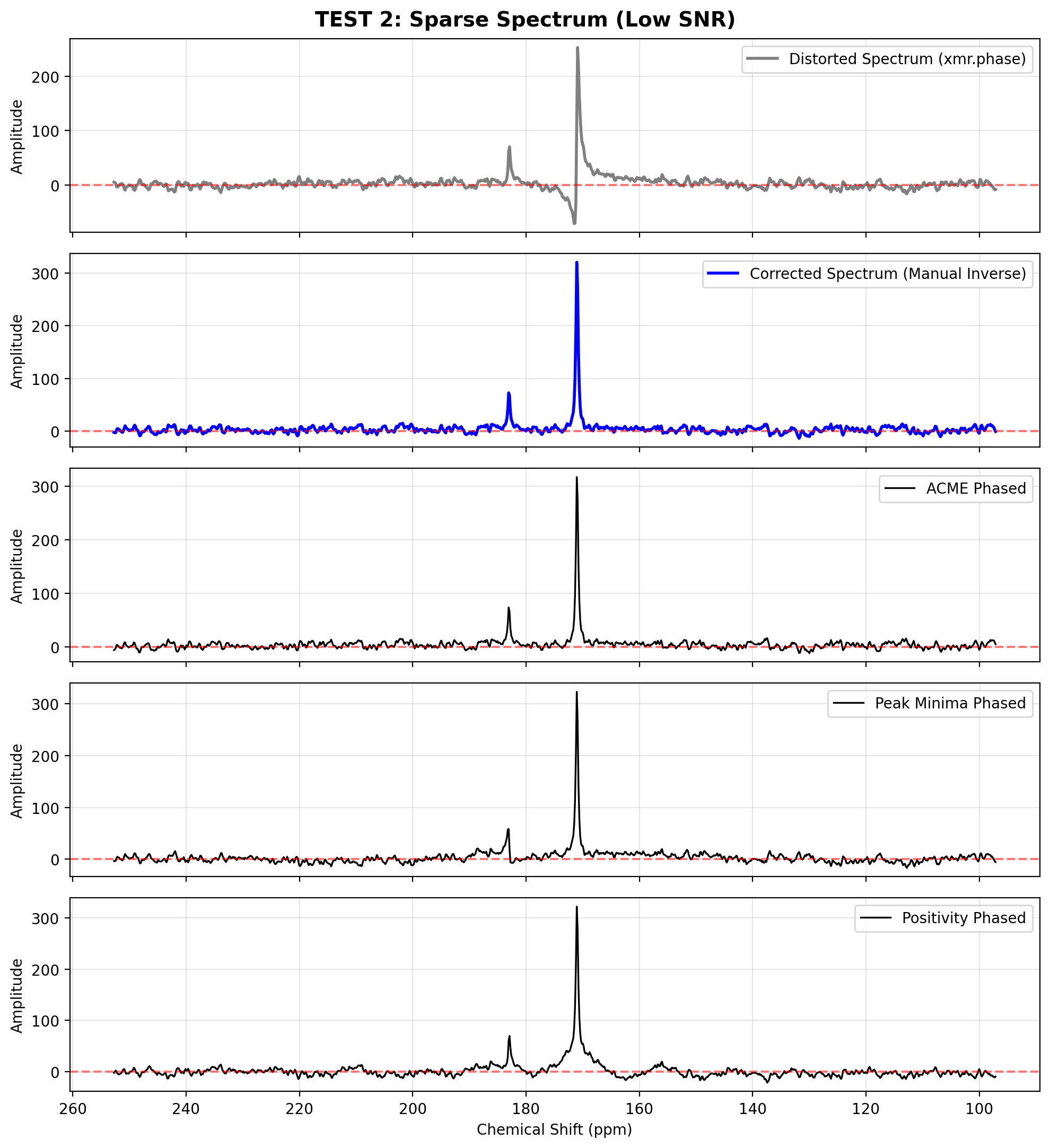
3. Ultra-Low SNR & Parameter Locking¶
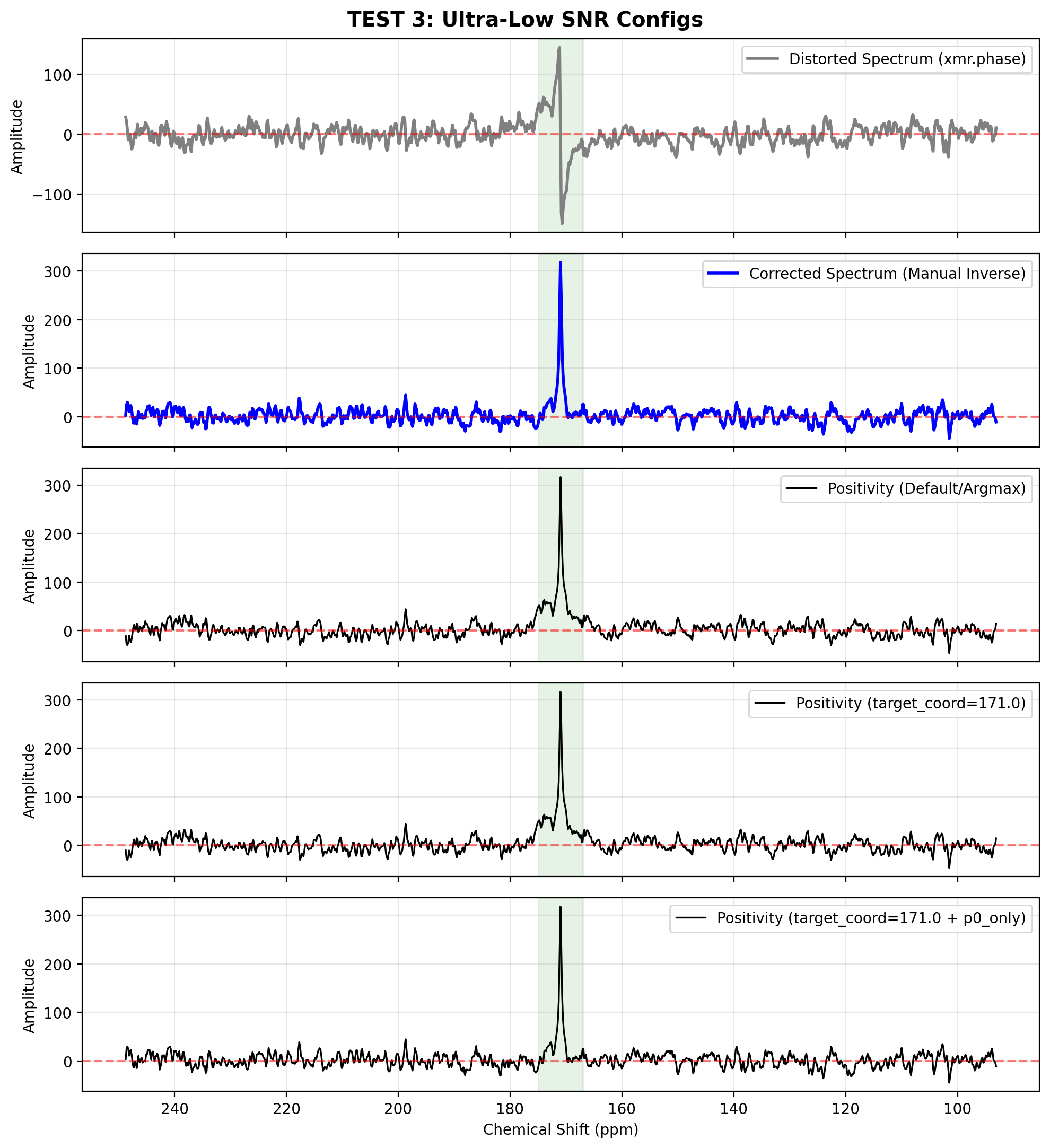
In extremely challenging SNR environments, fitting both and simultaneously can lead to the optimizer escaping into local minima by applying massive, unrealistic first-order twists.
You can restrict the solver by locking the first-order phase (p0_only=True) and forcing it to evaluate an explicit coordinate (target_coord) rather than guessing the peak location based on the absolute maximum (which might just be a noise spike).
Make another dummy 13C spectrum (even lower SNR)
# 1. Simulate an ideal, ULTRA-NOISY FID
fid_ultra = simulate_fid(
amplitudes=[100], chemical_shifts=[171.0],
reference_frequency=32.1, carrier_ppm=171.0, spectral_width=5000,
dampings=[15], dead_time=0.0, phases=0.0, target_snr=1.5, n_points=1024
)
spec_ultra = fid_ultra.xmr.apodize_exp(lb=10.0).xmr.to_spectrum().xmr.to_ppm()
# 2. Distort it (Heavy p0 twist, no p1 roll for this test)
p0_distort_u, p1_distort_u, pivot_u = 90.0, 0.0, 171.0
spec_distorted_u = spec_ultra.xmr.phase(
dim="chemical_shift", p0=p0_distort_u, p1=p1_distort_u, pivot=pivot_u
)
spec_corrected_u = spec_distorted_u.xmr.phase(
dim="chemical_shift", p0=-p0_distort_u, p1=-p1_distort_u, pivot=pivot_u
)# Step 1-3.
# --> Simulation of the spectrum is hidden in the collpased cell above.
# 4. Autophase (Testing Positivity configurations)
target = 171.0
roi_width = 8.0
pos_def = spec_distorted_u.xmr.autophase(
dim="chemical_shift", method="positivity", peak_width=roi_width
)
pos_tgt = spec_distorted_u.xmr.autophase(
dim="chemical_shift", method="positivity", peak_width=roi_width, target_coord=target
)
# Locking p1 to 0.0 stabilizes the solver in extreme noise
pos_tgt_p0 = spec_distorted_u.xmr.autophase(
dim="chemical_shift", method="positivity", peak_width=roi_width, target_coord=target, p0_only=True
)
plot_spectra(
[
(spec_distorted_u, "Distorted Spectrum (xmr.phase)"),
(spec_corrected_u, "Corrected Spectrum (Manual Inverse)"),
(pos_def, "Positivity (Default/Argmax)"),
(pos_tgt, f"Positivity (target_coord={target})"),
(pos_tgt_p0, f"Positivity (target_coord={target} + p0_only)")
],
"TEST 3: Ultra-Low SNR Configs",
target_coord=target,
peak_width=roi_width
)
4. Multi-Dimensional Datasets (mode)¶
When processing multi-dimensional data (such as kinetic series, dynamic nuclear polarization buildups, or repeated acquisitions), attempting to autophase every slice individually can be risky—especially for early time points where the signal is buried in noise.
The autophase method handles N-dimensional data natively via the mode parameter:
mode="single"(default): The algorithm searches the entire dataset to find the single 1D slice with the highest absolute magnitude. It runs the optimization strictly on that high-SNR slice, and then broadcasts the resulting and parameters across the entire dataset. This ensures perfectly consistent phase across your kinetic or repetition dimension.mode="all": (Coming Soon) Will independently calculate and apply unique phase parameters for every 1D slice.
Make a dummy 2D dataset (e.g., an amplitude buildup)
# 1. Simulate a 2D dataset (e.g., a buildup curve over 5 repetitions)
fids_2d = []
for amp in np.linspace(10, 100, 5):
fid = simulate_fid(
amplitudes=[amp, amp * 0.4], chemical_shifts=[171.0, 183.0],
reference_frequency=32.1, carrier_ppm=175.0, spectral_width=5000,
dampings=[15, 15], dead_time=0.0, phases=0.0, target_snr=15 / (100 - amp + 0.01) * 10, n_points=1024
)
fids_2d.append(fid)
# Stack into a 2D DataArray: (repetitions, time)
fid_2d = xr.concat(fids_2d, dim=xr.DataArray(np.arange(5), dims="repetitions", name="repetitions"))
spec_2d = fid_2d.xmr.apodize_exp(lb=10.0).xmr.to_spectrum().xmr.to_ppm()
# 2. Distort the entire dataset with a unified phase twist
p0_distort_m, p1_distort_m, pivot_m = -110.0, 450.0, 175.0
spec_distorted_2d = spec_2d.xmr.phase(
dim="chemical_shift", p0=p0_distort_m, p1=p1_distort_m, pivot=pivot_m
)# Step 1-2.
# --> Simulation of the 2D dataset is hidden in the collpased cell above.
# 3. Autophase the 2D dataset using mode="single"
# It will automatically find the slice with the highest signal (repetition 4)
# optimize the phase on that slice, and apply it globally.
spec_phased_2d = spec_distorted_2d.xmr.autophase(
dim="chemical_shift", method="positivity", peak_width=10.0, mode="single"
)
# 4. Plot the results across different slices to prove broadcasting worked
plot_spectra([
(spec_distorted_2d.isel(repetitions=-1), "Distorted (Max Signal Slice)"),
(spec_phased_2d.isel(repetitions=-1), "Phased (Max Signal Slice)"),
(spec_phased_2d.isel(repetitions=0), "Phased (Low Signal Slice, Broadcasted)")
], "TEST 4: Multi-Dimensional Data (mode='single')")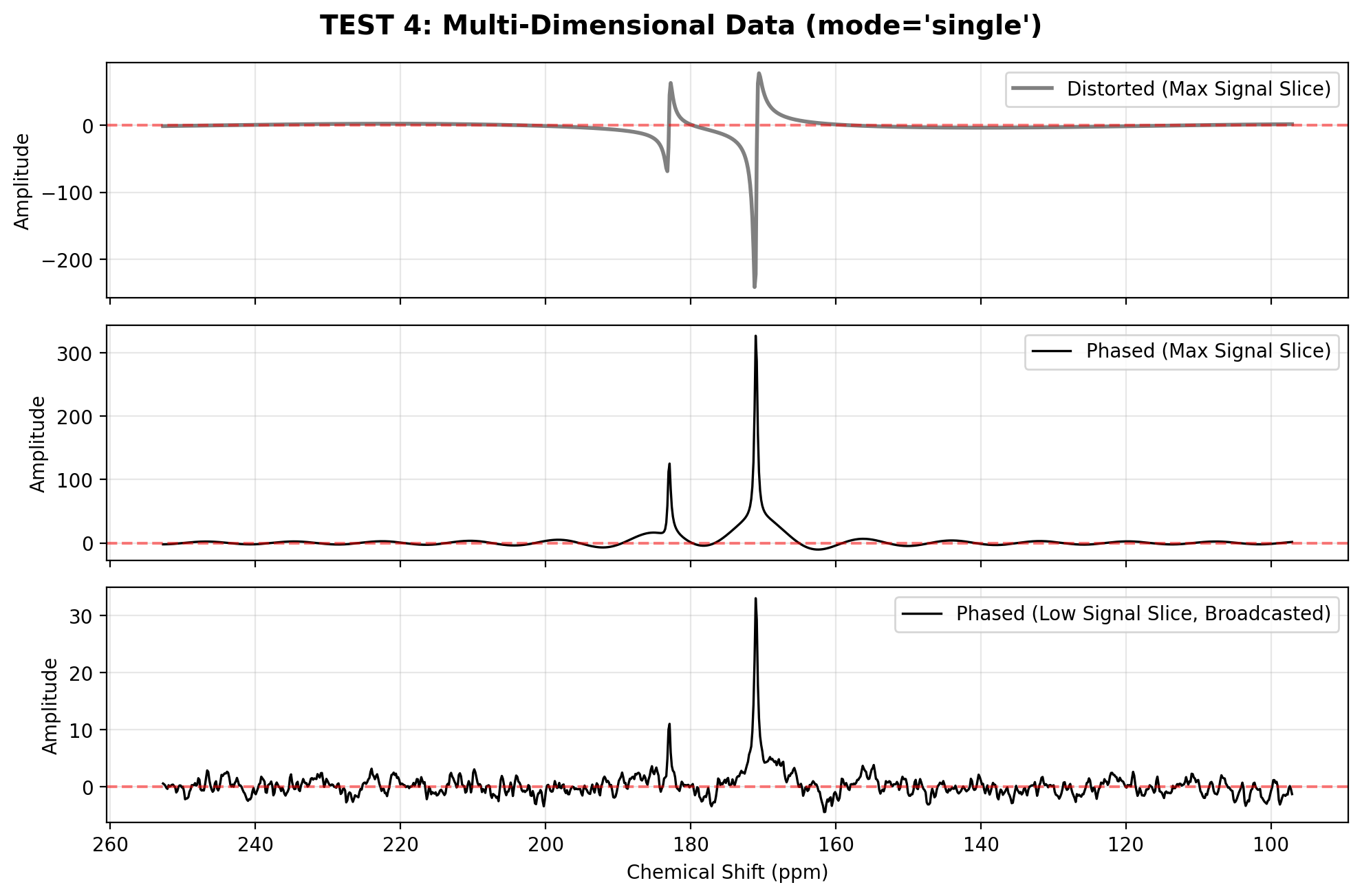
- Chen, L., Weng, Z., Goh, L., & Garland, M. (2002). An efficient algorithm for automatic phase correction of NMR spectra based on entropy minimization. Journal of Magnetic Resonance, 158(1–2), 164–168. 10.1016/s1090-7807(02)00069-1