Waterfall plots (also known as ridge or joy plots) are essential in Magnetic Resonance Spectroscopy (MRS) for visualizing kinetic, dynamic, or relaxation series. By vertically stacking and horizontally offsetting 1D spectra, they allow the human eye to easily track peak growth, decay, and frequency shifts over an independent variable like time .
In xmris, this visualization is built entirely around our WaterfallConfig object, giving you publication-ready results without cluttering your function calls.
1. Data Requirements¶
Before plotting, your xarray.DataArray must meet the following criteria:
Dimensionality: Must be at least 2D.
X-Axis (Horizontal): The accessor will automatically search for dimensions named
chemical_shiftorfrequency. If your spectral axis has a different name, you must specify it explicitly via thex_dimargument.Stack Axis (Vertical): The accessor will automatically stack along the remaining dimension. If multiple dimensions remain, it looks for
averagesorrepetitions, or you can specify it explicitly via thestack_dimargument.
2. Generating Synthetic Data¶
We will generate a realistic kinetic time-course of a hyperpolarized 13C experiment where Pyruvate decays and Lactate grows over 60 seconds.
Time series data generation
We will use the xmris.fitting.simulation module:
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from xmris.fitting.simulation import simulate_fid
from xmris.visualization import WaterfallConfig
# Define our time series
time_points = np.linspace(0, 60, 25)
fids = []
# Simulate the kinetic evolution
for t in time_points:
pyr_amp = 1000.0 * np.exp(-t / 20.0)
lac_amp = 400.0 * (1.0 - np.exp(-t / 20.0))
fid = simulate_fid(
amplitudes=[lac_amp, pyr_amp],
chemical_shifts=[183.3, 171.0],
reference_frequency=32.1,
carrier_ppm=171.0,
spectral_width=5000.0,
n_points=1024,
dampings=[10.0, 12.0],
phases=[0.0, 0.0],
lineshape_g=[0.0, 0.0],
target_snr=25.0
)
fids.append(fid)
# Combine into a 2D array by stacking along a new 'kinetic_time' dimension
da_kinetic_fid = xr.concat(fids, dim="kinetic_time").assign_coords(kinetic_time=time_points)
da_kinetic_fid.coords["kinetic_time"].attrs["units"] = "s"
# Convert to frequency-domain spectrum (ppm) and extract the real part
da_kinetic = da_kinetic_fid.xmr.to_spectrum().xmr.to_ppm().real3. Basic Usage¶
Because we utilize an intelligent xarray accessor, the simplest plotting call requires zero arguments. The accessor dynamically reads the dataset’s units and dimensions to build the axes automatically.
# We slice the region of interest, and the accessor handles the rest
ax = da_kinetic.sel(chemical_shift=slice(160, 190)).xmr.plot.waterfall()
plt.show()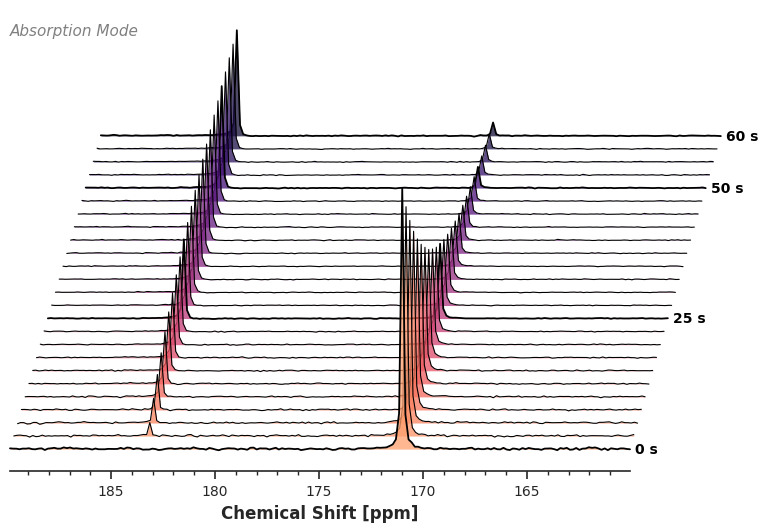
4. Advanced Configuration¶
To customize the plot, do not pass endless keyword arguments. Instead, instantiate a WaterfallConfig. Outputting this object in a notebook renders a table of all available styling options.
CFG = WaterfallConfig()
CFGYou can update these parameters using standard attribute assignment. Let’s create a dynamic, pseudo-3D angled sheer with heavy overlap.
CFG.stack_scale = 10.0
CFG.stack_offset = 0.5
CFG.stack_skew = -20.0
CFG.cmap = "viridis"
CFG.annotation = None
CFG.xlabel = r"$^{13}\mathrm{C}$ Chemical Shift"
# Pass the config to the accessor
ax = da_kinetic.sel(chemical_shift=slice(160, 190)).xmr.plot.waterfall(config=CFG)
plt.show()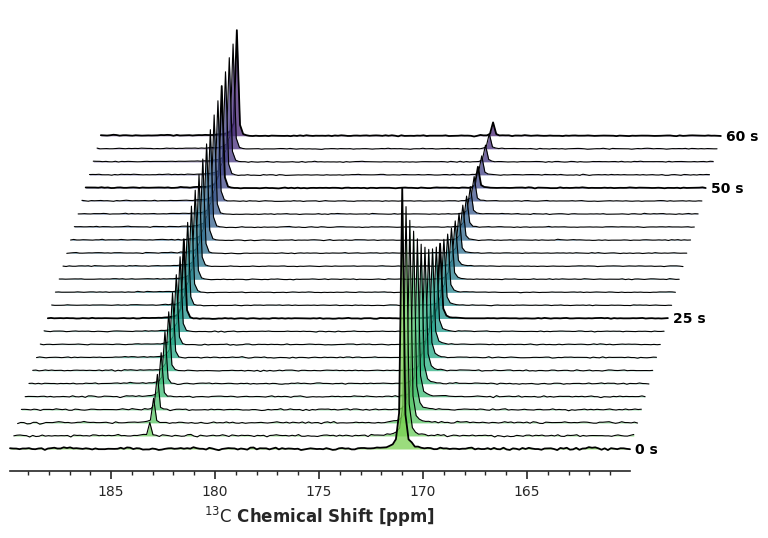
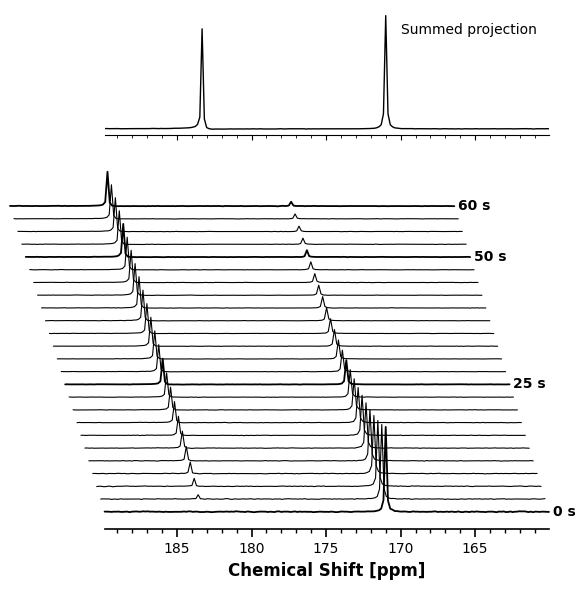
5. Subplot Integration (Wireframe Mode)¶
Professional library functions should never hijack your global plotting environment. Because plot.waterfall() returns a standard matplotlib.Axes object and accepts an ax keyword argument, you can easily embed these visualizations into multi-panel figures.
Here, we also demonstrate the Wireframe Mode by setting cmap=None, which disables the filled polygons for a clean, minimalist look.
from matplotlib.offsetbox import AnchoredText
# Configure a wireframe plot
CFG_WIRE = WaterfallConfig(
cmap=None, # Disables fill_between
annotation=None,
stack_scale=10.0,
stack_offset=1.5,
stack_skew=10.0
)
# Create a custom multi-panel figure
fig, (ax_top, ax_bottom) = plt.subplots(
nrows=2,
figsize=(6, 6),
gridspec_kw={"height_ratios": [1, 3]},
sharex=True,
)
# 1. Plot a standard 1D summed spectrum on the top panel
da_kinetic.sel(chemical_shift=slice(160, 190)).sum(dim="kinetic_time").plot.line(ax=ax_top, color="black", linewidth=1)
text_box = AnchoredText("Summed projection", loc="upper right", frameon=False, prop=dict(fontsize=10))
ax_top.add_artist(text_box)
ax_top.invert_xaxis()
ax_top.set_xlabel("")
ax_top.set_ylabel("")
ax_top.set_yticks([])
for spine in ["left", "right", "top"]:
ax_top.spines[spine].set_visible(False)
# 2. Inject the waterfall plot into the bottom panel
_ = da_kinetic.sel(chemical_shift=slice(160, 190)).xmr.plot.waterfall(ax=ax_bottom, config=CFG_WIRE)
plt.tight_layout()
plt.show()