Handling Complex MRI/MRS Data¶
Magnetic resonance data is inherently complex-valued, but many standard tools lack native support for complex numbers. Splitting this data into separate real and imaginary channels is frequently required for:
Machine Learning:
Routing data through standard PyTorch or TensorFlow architectures.
Serialization:
Saving
xarraydatasets to disk, e.g.da.to_netcdf()andxr.open_dataset().Visualization:
Plotting real and imaginary components independently.
The xmris accessor provides dedicated utilities (to_real_imag and to_complex) to safely reshape and reconstruct complex xarray.DataArray objects while preserving all metadata and physical coordinates.
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
# Ensure the accessor is registered
import xmris1. Splitting Complex Data into Channels¶
Let’s generate a synthetic complex Free Induction Decay (FID) signal.
# Generate a synthetic complex FID
t = np.linspace(0, 1, 512)
complex_fid = np.exp(-t * 3.0) * np.exp(1j * 2 * np.pi * 15.0 * t)
da_complex = xr.DataArray(
complex_fid,
dims=["time"],
coords={"time": t},
attrs={"B0": 3.0},
name="Signal",
)
print("Original Shape:", da_complex.shape)
print("Original Dtype:", da_complex.dtype)Original Shape: (512,)
Original Dtype: complex128
By calling .xmr.to_real_imag(), the array is expanded with a new dimension containing the real and imaginary components.
da_split = da_complex.xmr.to_real_imag()
print("\nSplit Shape:", da_split.shape)
print("Split Dtype:", da_split.dtype)
da_split
Split Shape: (512, 2)
Split Dtype: float64
The array cleanly expanded from (512,) to (512, 2). Because it remains a standard xarray.DataArray, we can easily plot the two channels side-by-side.
fig, ax = plt.subplots(figsize=(8, 4))
da_split.plot.line(ax=ax, x="time", hue="component")
ax.set_title("FID Split into Real and Imaginary Components")
plt.show()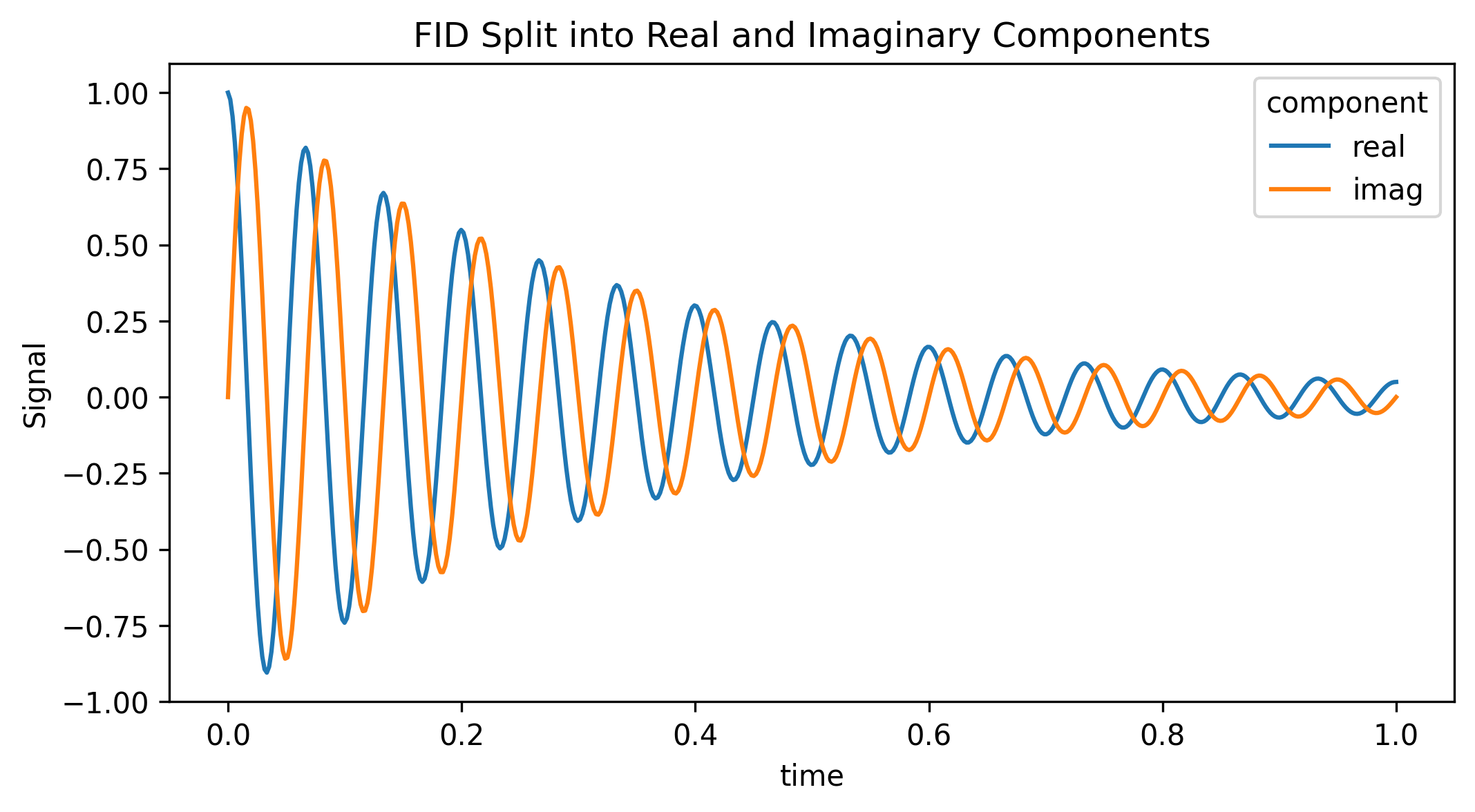
2. Customizing the Split Dimension¶
If your specific ML pipeline expects the channel dimension to have a specific name (e.g., "channel" instead of "component"), you can pass these arguments directly to the function.
da_torch = da_complex.xmr.to_real_imag(dim="channel", coords=("ch0", "ch1"))
print("Custom Dimension:", da_torch.dims)Custom Dimension: ('time', 'channel')
3. Reconstructing the Complex Array¶
After processing the split channels (e.g., passing them through a neural network), you can seamlessly collapse the dimension back down to a standard complex array for downstream signal processing (like an FFT) using .xmr.to_complex().
da_recon = da_split.xmr.to_complex()
is_identical = np.allclose(da_complex.values, da_recon.values)
print(f"Shape Restored: {da_recon.shape}")
print(f"Data Type Restored: {da_recon.dtype}")
print(f"Exact Mathematical Recovery: {is_identical}")Shape Restored: (512,)
Data Type Restored: complex128
Exact Mathematical Recovery: True